The discovery of a tRNA modification enzyme that also acts on nucleosides
Elucidation of the substrate recognition mechanism of ArcS, the second-step enzyme for archaeosine synthesis in tRNA
Previously reported tRNA modification enzymes act only on RNA polymers. However, it has been discovered that ArcS, the second-step enzyme for archaeosine synthesis, possesses unprecedented activity by acting even on a nucleoside.
The genetic information on DNA is transcribed into messenger RNA (mRNA) and translated to the amino acid sequence by transfer RNA (tRNA) on the ribosome. Modified nucleosides within RNA are involved in maintaining and regulating the protein synthesis system. Archaeosine is a modified nucleoside found only in the tRNAs from archaea, the so-called third domain of life, and contributes to the maintenance of the L-shaped tRNA three-dimensional structure. The synthesis of archaeosine involves multiple steps, with the first step introducing a preQ0 base into tRNA via ArcTGT. In the second step, ArcS transfers an amino acid, lysine, to the preQ0 base in tRNA and synthesizes preQ0-Lys as an intermediate. The resultant preQ0-Lys in tRNA is then converted into archaeosine by RaSEA, the third-step enzyme.
This synthesis pathway of archaeosine was elucidated in 2019 through a collaborative study by Ehime University and Gifu University (Figure 1: Yokogawa et al., Nature Chem. Biol. (2019)). However, the substrate specificity of the second-step enzyme ArcS was previously unknown.
To address this issue, a research group led by Professor Hiroyuki Hori, Lecturer Dr. Ryota Yamagami, and graduate students Shu Fujita, Yuzuru Sugio, and Dr. Takuya Kawamura (currently at Thomas Jefferson University, USA) at the Graduate School of Science and Engineering, Ehime University, in collaboration with Professors Takashi Yokogawa and Natsuhisa Oka from Gifu University and Associate Professor Akira Hirata from Tokushima University, conducted biochemical analyses.
Most RNA modification enzymes recognize the three-dimensional structure around the target site of RNA and only rarely the RNA sequence itself. To investigate the substrate RNA specificity of ArcS, preQ0-modified tRNA was fragmented using DNAzymes, and lysine transfer was assessed for each fragment (Figure 2). Surprisingly, ArcS transferred lysine to all RNA fragments containing preQ0. In the 21-nucleotide (21 nt) RNA fragment, not only the whole tRNA structure, but also the D-arm structure was disrupted. This result demonstrates that ArcS does not recognize the three-dimensional structure of substrate RNA. To identify the minimum substrate, lysine-transfer was assessed using the preQ0 base, preQ0 nucleoside, 5′-phosphorylated preQ0 nucleotide, and 3′-phosphorylated preQ0 nucleotide (Figure 3). It was found that the minimum substrate was the preQ0 nucleoside, with reaction efficiency increasing when a phosphate group was attached to the 5′ position. Thus, ArcS is an unprecedented tRNA modification enzyme that can act on a nucleoside as the substrate.
With the development of mRNA vaccines for COVID-19, modified nucleosides like pseudouridine and 1-methylpseudouridine are effectively used, and research on introducing various modifications into target RNAs are being conducted globally. The discovery of ArcS, which can utilize a nucleoside as a minimum substrate, provides new insights into the synthesis of precursor molecules for these modified nucleosides.
The results of this research were published online in the Journal of Biological Chemistry on June 27, 2024.
Reference URL: https://www.jbc.org/article/S0021-9258(24)02006-4/fulltext
Bibliographic Information
“ArcS from Thermococcus kodakarensis transfers L-lysine to preQ0 nucleoside derivatives as minimum substrate RNAs.”,
Shu Fujita, Yuzuru Sugio, Takuya Kawamura, Ryota Yamagami, Natsuhisa Oka, Akira Hirata, Takashi Yokogawa, and Hiroyuki Hori,
Journal of Biological Chemistry, June 27, 2024.
doi: 10.1016/j.jbc.2024.107505.
Fundings
- This work was partly supported by Grant-in-Aid for Scientific Research from the JAPAN Society for the Promotion of Science (JSPS) KAKENHI Grant Numbers JP20H03211 and JP24K09381 and Funding from the Institute for Fermentation, Osaka, Grant Number G-2022-2-052.
Media
-
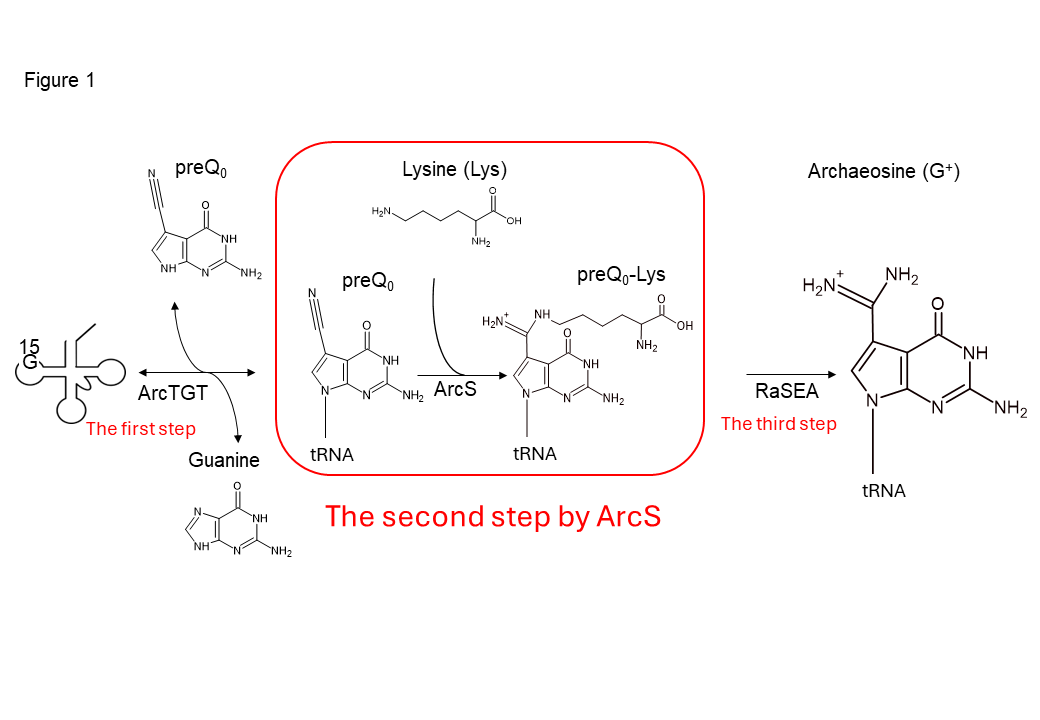
Figure 1 Synthesis of archaeosine in tRNA
Archaeosine is synthesized via three steps in tRNA. ArcS, the second-step enzyme, transfers a lysine to preQ0, which is introduced into tRNA by ArcTGT, the first-step enzyme. The resultant preQ0-Lys in tRNA is further converted to archaeosine by RaSEA, the third-step enzyme. In this study, the enzymatic properties of ArcS have been investigated.
credit : Hiroyuki Hori, Shu Fujita, Ehime University
Usage Restriction : Please get copyright permission -
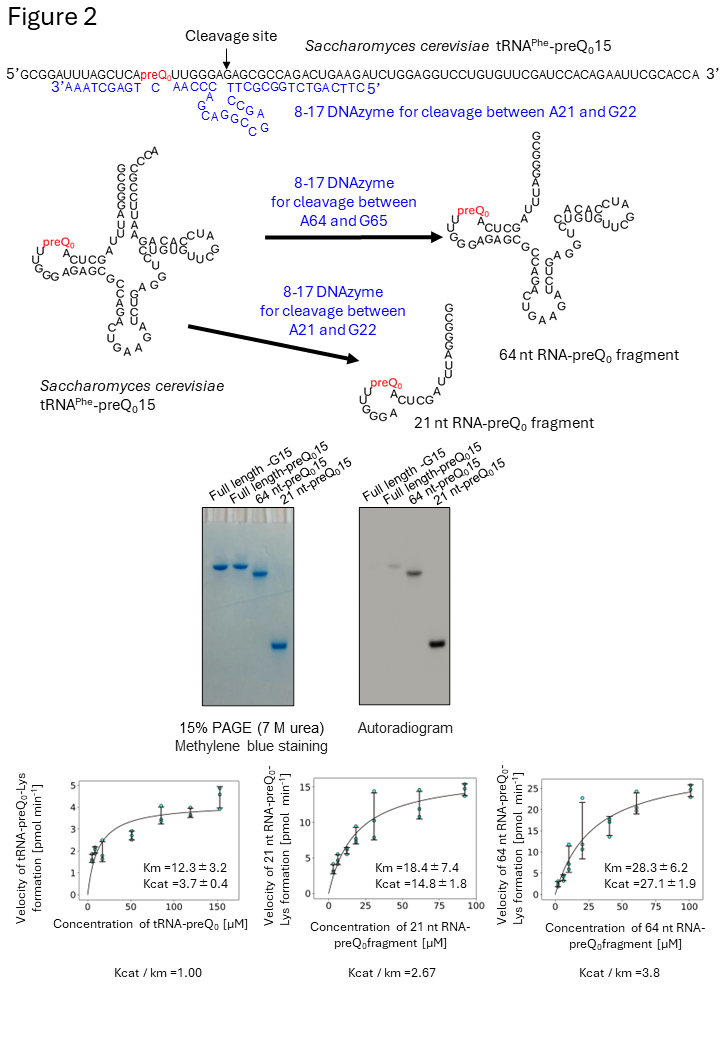
Figure 2 ArcS does not recognize the three-dimensional structure of substate RNA.
To clarify the recognition site(s) in tRNA, tRNA-preQ0, which is highlighted in red, was individually digested by DNAzymes (blue) and two RNA-preQ0 fragments (64 nt and 21 nt) were prepared. To our surprise, 14C-labeld lysine was transferred to all RNAs, which contain preQ0. This result demonstrates that ArcS does not recognize the three-dimensional structure of substate RNA.
credit : Hiroyuki Hori, Shu Fujita, Ehime University
Usage Restriction : Please get copyright permission -
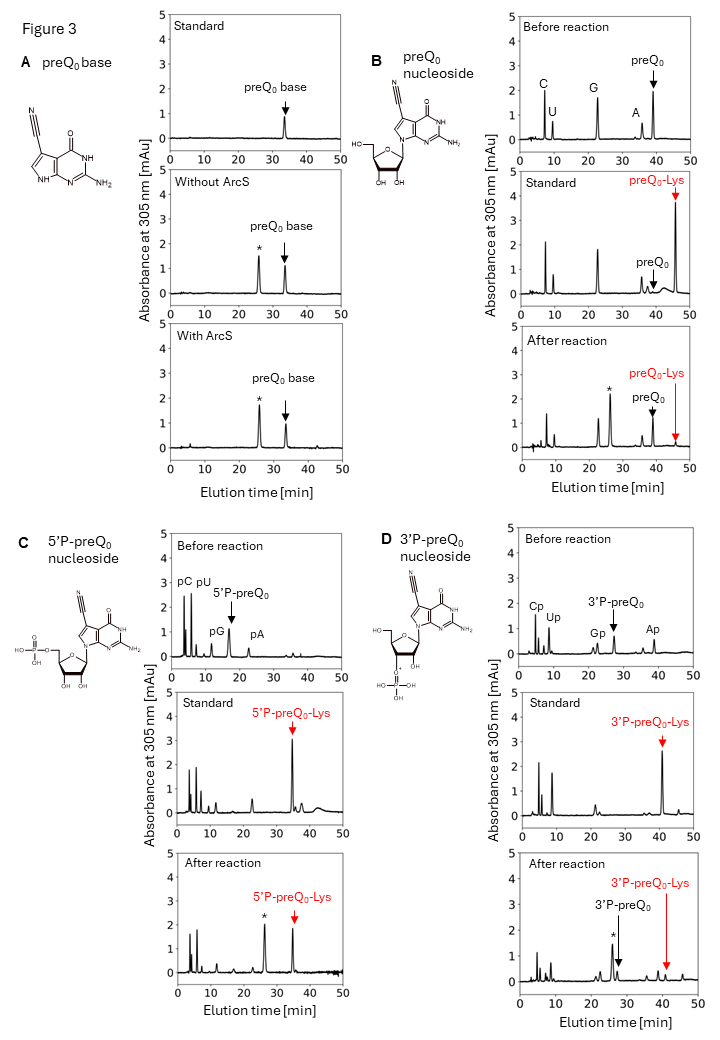
Figure 3 the minimum substrate of ArcS is preQ0 nucleoside.
To clarify the minimum substrate of ArcS, preQ0 base, preQ0 nucleoside, 5’-phosphate preQ0 nucleotide, and 3’-phosphate preQ0 nucleotide were tested. Although lysine was not transferred to a pre Q0 base, lysine-transfer was observed into preQ0 nucleoside, 5’-phosphate preQ0 nucleotide, and 3’-phosphate preQ0 nucleotide. Thus, the minimum substrate of ArcS is preQ0 nucleoside.
credit : Hiroyuki Hori, Shu Fujita, Ehime University
Usage Restriction : Please get copyright permission
Contact Person
Name : Hiroyuki Hori
Phone : +81-89-927-8548
E-mail : hori.hiroyuki.my@ehime-u.ac.jp
Affiliation : Graduate School of Science and Engineering, Ehime University
