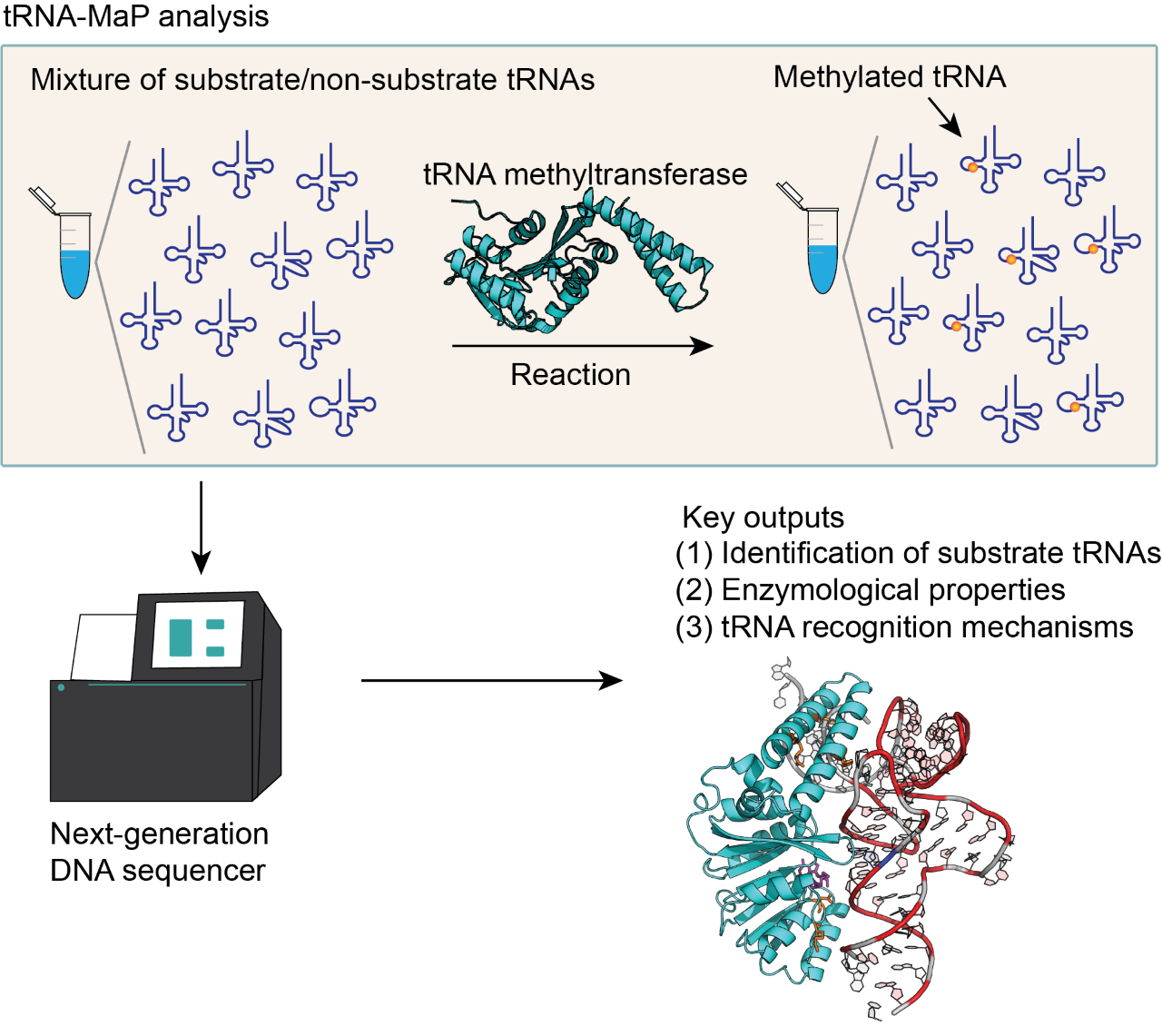
tRNA-MaP: functional analyses of RNA-related enzymes using a next-generation DNA sequencer
tRNA-MaP can rapidly analyze the specificity of tRNA m1A22 methyltransferase (TrmK) for substrate tRNAs.
We have developed a new method, tRNA-Map, that can analyze RNA-related enzymes using next-generation DNA sequencing. We analyzed tRNA m1A22 methyltransferase (TrmK) using tRNA-Map. Mutation profiling revealed that TrmK selects a subset of tRNAs for the substrate. Using 240 variants of tRNALeu transcripts, we found that U8, A14, G15, G18, G19, U55, Purine57 and A58 are important for the methylation by TrmK. In addition, a docking model between TrmK and tRNA has been constructed.
Genetic information encoded in genomic DNA is transcribed to mRNAs and then the codons on mRNA are decoded by transfer RNAs (tRNAs) during protein synthesis. tRNAs deliver amino acids to ribosomes and proteins are synthesized from the amino acids on the ribosomes according to the decoded genetic information. Therefore, tRNA plays a key role during the translation of genetic information. tRNAs contain numerous modified nucleosides, which regulate the accuracy and efficiency of protein synthesis. Modified nucleosides in tRNA are synthesized by tRNA modification enzymes. Therefore, unveiling the mechanisms by which tRNA modification enzymes selectively recognize substrate tRNAs from non-substrate RNAs; the when, where, and how many tRNAs are being modified by the modification enzymes, is of crucial importance to understand the protein synthesis machinery. Addressing these key questions is, however, challenging due to the lack of a high-throughput technique that identifies the characteristic properties of tRNA modification enzymes.
To overcome this issue, Drs. Yamagami and Hori at Ehime University applied next-generation DNA sequencing technology to functional analyses of tRNA modification enzymes and developed a new high-throughput assay method, “tRNA-MaP”. The tRNA-MaP technique can rapidly screen an RNA pool consisting of more than 5,000 RNA species and identify the substrate tRNAs of the target tRNA modification enzyme(s) with comparative sensitivity to already-established methods. By tRNA-MaP, in combination with protein orthology analyses, we predicted numerous natural modifications in Geobacillus stearothermophilus tRNAs. Furthermore, we analyzed the substrate recognition mechanism of G. stearothermophilus tRNA m1A22 methyltransferase (TrmK), which methylates adenosine at position 22 to 1-methyladenosine (m1A22) in tRNA, using tRNA-Map. Mutation profiling has revealed that TrmK selects a subset of tRNAs for the substrate. Using 240 variants of G. stearothermophilus tRNALeu transcripts, we found that U8, A14, G15, G18, G19, U55, Purine57 and A58 are important for the methylation by TrmK. In addition, based on the recognition sites in tRNA and the crystal structure of TrmK, a docking model between TrmK and tRNA has been constructed.
This study has revealed that tRNA-Map is applicable for the analysis of the tRNA modification enzyme. Notably, because tRNA-Map can analyze any RNA molecular species from any organism, even DNA molecules, tRNA-Map can be used for analysis of all nucleic acid-related proteins except for tRNA modification enzymes. Thus, tRNA-Map can accelerate the integrative understanding of the flow of genetic information.
Reference URL: https://www.jbc.org/article/S0021-9258(22)01202-9/fulltext#relatedArticles
Bibliographic Information
Application of mutational profiling: new functional analyses reveal the tRNA recognition mechanism of tRNA m1A22 methyltransferase (TrmK), Ryota Yamagami and Hiroyuki Hori, Journal of Biological Chemistry, in press. DOI:https://doi.org/10.1016/j.jbc.2022.102759
Fundings
- JSPS KAKENHI Grant Number JP22K15035,JP21K20638,JP20H03211
- Ehime University Grant Number ZI07b
Media
Contact Person
Name : Ryota Yamagami / Hiroyuki Hori
Phone : +81-89-927-9919 / +81-89-927-8548
E-mail : yamagami.ryota.bn@ehime-u.ac.jp, hori.hiroyuki.my@ehime-u.ac.jp
Affiliation : Graduate School of Science and Engineering